siDirectPlus
 Efficient and Secure siRNA Design Tool
Efficient and Secure siRNA Design Tool– Now Available Offline!
siDirectPlus 2024 Edition Now Available! ・ A new feature has been added: allows gene extraction with sequences complementary to the seed region in the 3'-UTR region as off-target candidate genes! ・ Now you can input gene sequences by referencing the bundled database enabling completely offline use! |
Overview
siDirect is a sequence design tool for siRNA required to induce RNAi in mammalian cells. A major advantage of siDirect is its ability to design sequences that suppress off-target effects. Since the first version was released in 2004, researchers have been able to access the website to design siRNAs.
siDirectPlus is the offline version of siDirect. Sequence information explored in drug discovery is highly confidential data that cannot be handled online. Therefore, we recommend using siDirect Plus, which offers equivalent functionality offline. Off-target searches can be integrated with the GGGenome high-speed base search engine; an offline package for this is also available.
siDirectPlus is the offline version of siDirect. Sequence information explored in drug discovery is highly confidential data that cannot be handled online. Therefore, we recommend using siDirect Plus, which offers equivalent functionality offline. Off-target searches can be integrated with the GGGenome high-speed base search engine; an offline package for this is also available.
Key Features
 |
1. Designs and produces highly effective siRNAs siDirectPlus uses patented technology (JPO3839453) proposed by Associate Professor K Ui-Tei et al. Efficacy rate of the designed siRNAs was 98%. |
 |
2. Accurate Prediction for Safety Using the GGGenome search package for off-target effect prediction enables precise and comprehensive searches. Additionally, siRNA with low off-target effects can be selected based on the calculated parameters. |
 |
 |
| Reliable Services and Support | Custom Design Services |
| We provide know-how to support your product utilization, including considerations for siRNA design. | We also offer RNA molecule design services. We can assist when in-house design proves difficult at your site. |
Additional Features on Packaged Version
With siDirectPlus, you can check similar base sequences on huge genome database using GGGenome.
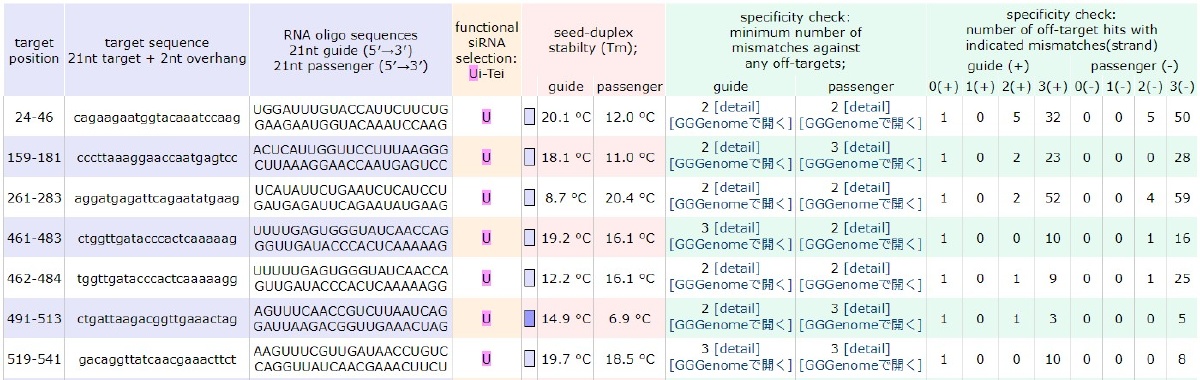
Similar base sequences can be confirmed using the GGGenome package version, allowing you to select the search database from the Drug Discovery Basic Package or adjust the number of mismatches for your search.
You can check the hit positions of off-target candidate genes.
You can check the hit positions of off-target candidate genes.
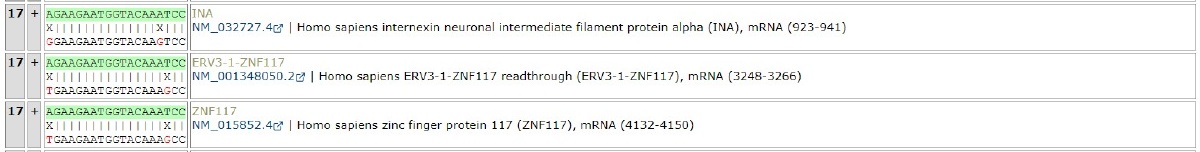
You can check the position where hits occur on off-target candidate genes.
Product Format
・ Provided on a USB flash drive or an SSD memory.
・ Provided as a set with the GGGenome package version (latest version).
・ Provided as a set with the GGGenome package version (latest version).
System Requirements
・ This product is compatible with 64-bit versions of Windows.
・ Windows 11 Pro version 21H2 or later, or Enterprise/Education version 21H2 or later
・ Windows 10 Pro 21H1 or later, or Enterprise/Education 20H2 or later
・ Hyper-V must be enabled (may be mutually exclusive with virtualization software like VMware)
・ Memory: 8GB or more (At least 4GB must be allocated to Docker for Windows)
・ Windows 11 Pro version 21H2 or later, or Enterprise/Education version 21H2 or later
・ Windows 10 Pro 21H1 or later, or Enterprise/Education 20H2 or later
・ Hyper-V must be enabled (may be mutually exclusive with virtualization software like VMware)
・ Memory: 8GB or more (At least 4GB must be allocated to Docker for Windows)









